About
This is a Comprehensive database for Deleterious synonymous variations prediction (CDsyn) that provides plentiful functional annotation in evaluating the importance of synonymous variations. With the assistance of CDsyn, our understanding of the role of synonymous mutations in human diseases is expected to improve.
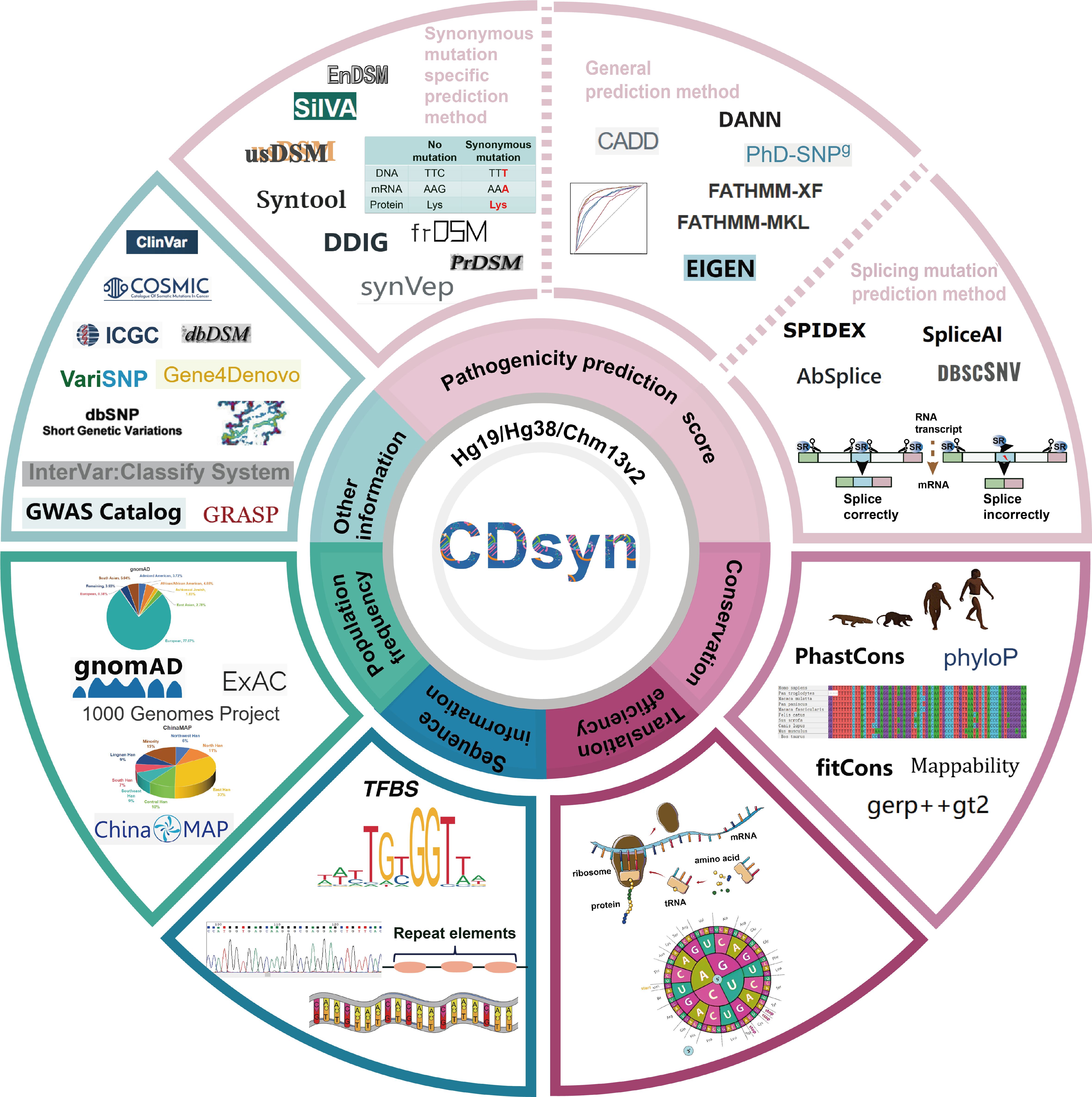
Figure 1. The data structure of CDsyn.
The CDsyn includes six-part contents, pathogenicity prediction scores (synonymous mutation specific pathogenicity prediction algorithms, general pathogenicity prediction algorithms and splicing mutation pathogenicity prediction algorithms), evolutionary conservations, sequence information, translation efficiency, and otherannotaion information.
The existing synonymous mutation related algorithms are comprised of CADD, DANN, fathmm-MKL, fathmm_xf, PhD-SNPg, DDIG, frDSM, EnDSM, usDSM, PrDSM, SilVA, and syntool. The first five tools are general prediction methods of deleterious variants and are used to predict the deleteriousness of synonymous mutations in our research. The rest tools are synonymous mutation specific pathogenicity prediction tools. Synvep, spidex, SpliceAI, AbSplice, and dbscSNV are splice mutation related algorithms
For the convenience and simplicity of browsing, we have filtered out some potentially more important information for online display. If you want to obtain complete information, please obtain the link to download the complete data on the "Download" page.
 Introduction
Introduction
 Search
Search
 Download
Download
 Help
Help